Theme 01
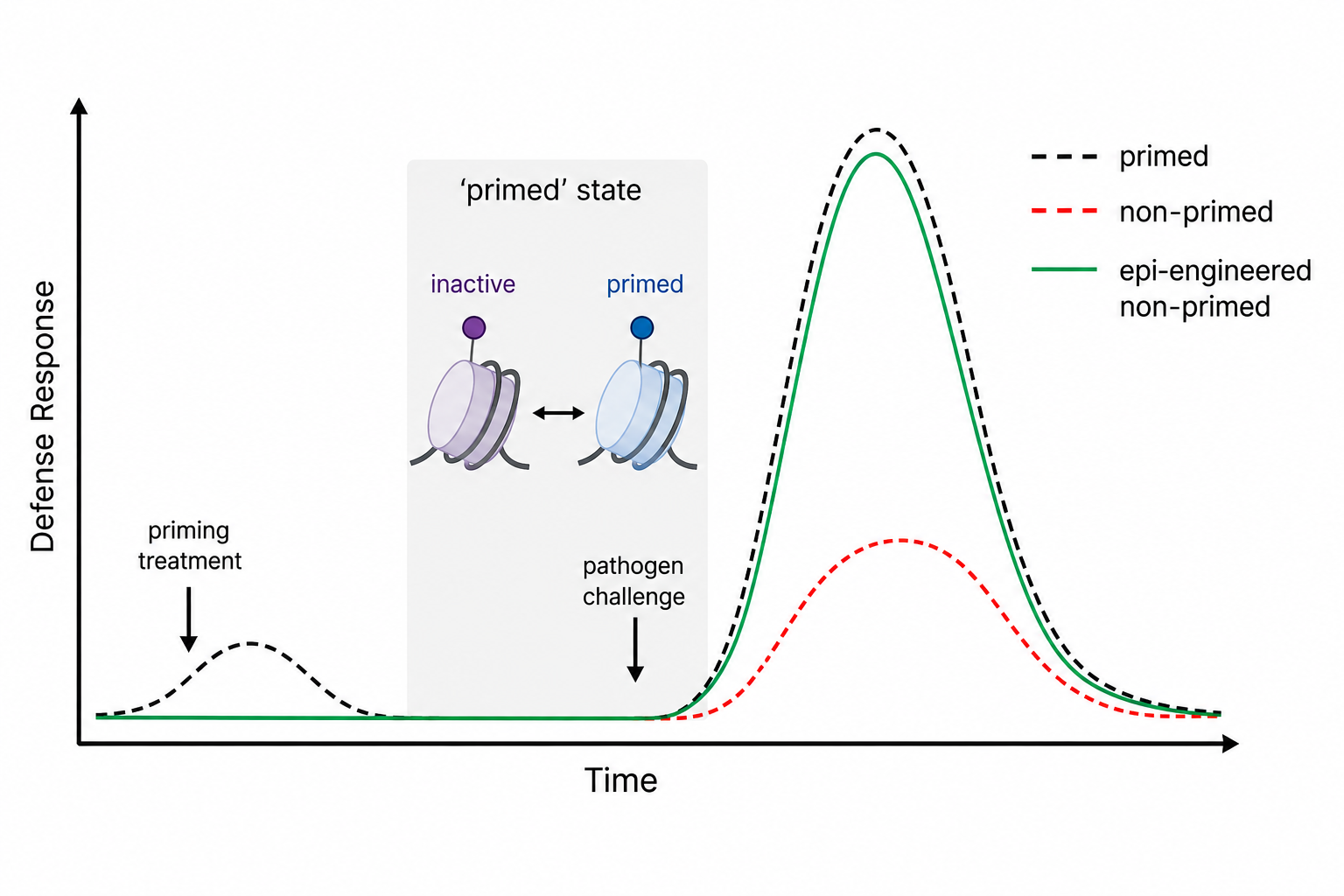
Priming Plants
Plants are particularly well equipped to respond rapidly, robustly and efficiently to stresses they have previously experienced. This transition to a 'primed' state of readiness demonstrates that plants form memories of life events — a phenomenon known to scientists for nearly a century. Where is this information stored? Every plant cell can encode information within chromatin, allowing the memory of life history to be stored and recalled in a decentralised manner across tissues. Our lab aims to decipher the chromatin code underlying transcriptional memory, particularly in the context of disease pressure. We combine epigenomics, bioinformatics, pathogen challenge, and epigenome engineering (see Theme 2) with the goal of rationally designing plants with enhanced resistance.
Theme 02
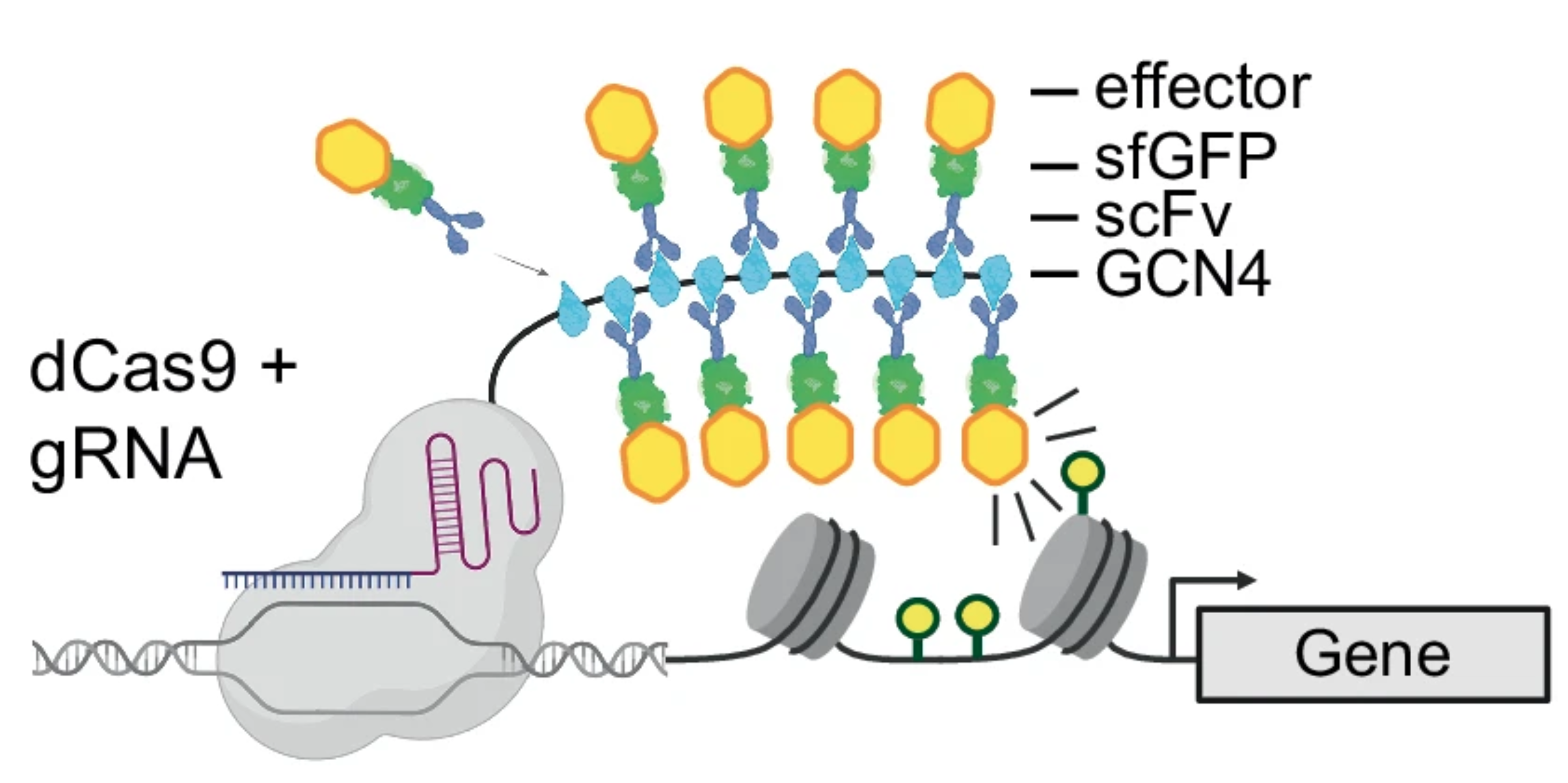
Epigenome Engineering
CRISPR has given us unprecedented access to the genome, and the same machinery can be repurposed to recruit chromatin-modifying activity to specific loci. This matters because one of the central challenges in chromatin biology is separating correlation from causation, and direct effects from indirect ones. Epigenome-engineering tools let us ask conceptually simple but fundamental questions — what happens when we put this mark here? Given the central role of chromatin in processes ranging from gene expression to ageing, these tools also open the door to non-GM crop improvement strategies with genomic precision. Yet the toolkit, and our understanding of the rules governing its efficacy, remains in its infancy. We are developing a suite of tools for modifying chromatin with spatial and temporal control — towards engineering chromatin with 'any mark, anytime, anywhere'.
Theme 03
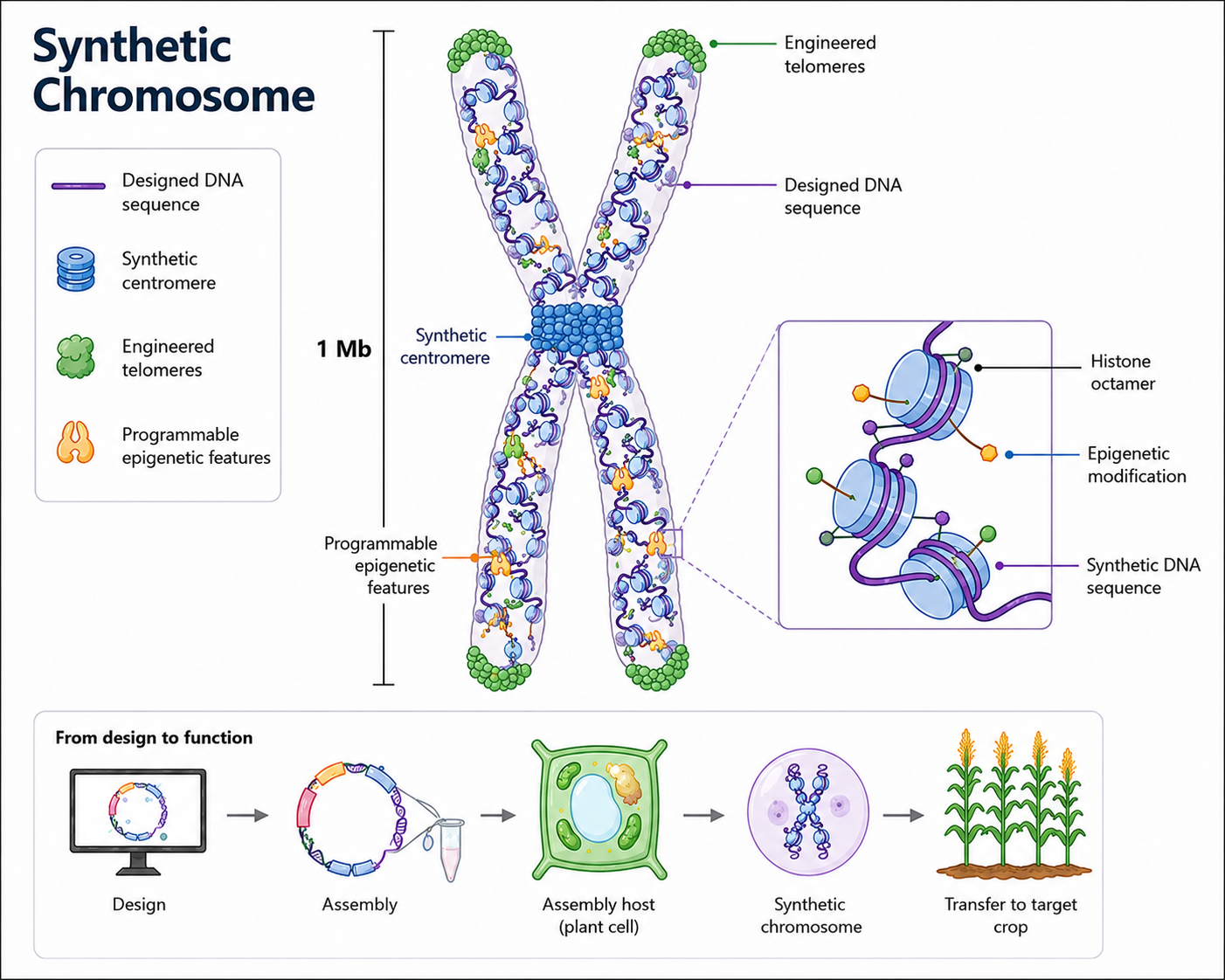
Synthetic Chromosomes
Traditional genetic modification lets us alter one or a handful of genes at a time. What if we could introduce tens, hundreds, even thousands at once? Entire complex traits — new spectrums of disease resistance, redesigned photosynthetic architectures, improved systems for nutrient uptake and drought tolerance — could be deployed in a single shot. Plant Artificial Chromosomes (PACs) hold the potential to make this possible. Through ARIA's Programmable Plants programme, we are working in a team along with biotechs, genome foundries, and leading plant synthetic biologists to turn PACs into a working technology. The project sits right at the edge of what is currently possible, alongside parallel efforts in other organisms — most notably the Sc2.0 yeast project and emerging work on synthetic human chromosomes. Advances in generative biology, functional genomics, centromere biology, and DNA synthesis are now converging to make the plant version tractable.
Theme 04
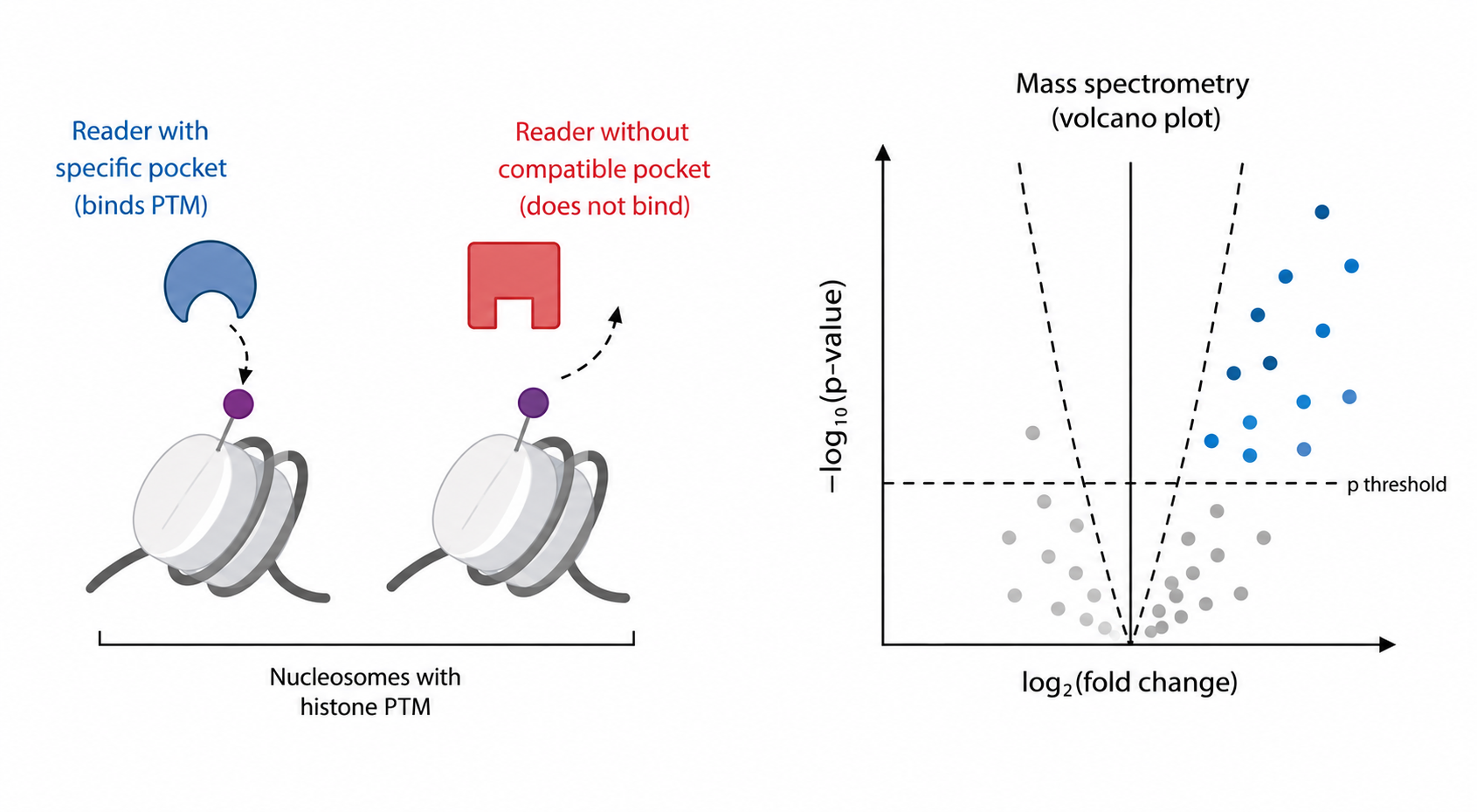
Chromatin Readers
The genome is decorated by a vast array of chromatin marks. These provide rich information about the function of the underlying sequence — distinguishing genes from transposons, coding regions from regulatory ones, and marking the chromosomal features required for mitotic and meiotic division. Yet our understanding of what these molecular signposts actually do lags far behind our ability to map them. A powerful way to reveal their function is to identify the cellular factors that perceive them, and then to dissect how those factors act. We use comparative interactomics — a quantitative mass-spectrometry-based approach developed with our collaborators — to identify the readers of key chromatin marks. Once a candidate is in hand, we can begin to uncover its function through a combination of genetics, genomics, biochemistry and computational analysis.
